Research Theme
Theoretical Studies on Dynamics behind Reactions, Functions, and Glass Transitions in Many-Body Molecular Systems
Keywords
Fluctuation, Reaction, Function, Glass transition
Many-body molecular systems, such as liquids and biomolecules, show complicated dynamics over a wide range of spatiotemporal scales and yield various thermodynamic properties. For example, in supercooled liquids, spatiotemporal non-uniform motions are found. The motions known as dynamic heterogeneity are now considered a crucial clue to understanding supercooled liquids and glass transition. The heterogeneous dynamics affect reaction dynamics. Furthermore, recent experimental and theoretical studies have demonstrated that reactions and conformational dynamics at single-molecule level are described by non-Poisson processes. In addition to these examples, it has been known that conformational changes with proper time scales are essential to protein functions. Thus, understanding heterogeneous dynamics in the many-body molecular systems is essential to elucidate thermodynamics and dynamic properties, reactions, and functions. So far, we have studied the complicated dynamics in these systems using multidimensional spectroscopy and multi-time correlation functions. Now, we are theoretically and computationally investigating how reactions proceed and biological functions are generated in fluctuating environments and how conformational dynamics yield interesting thermodynamic properties and change toward glass transition.
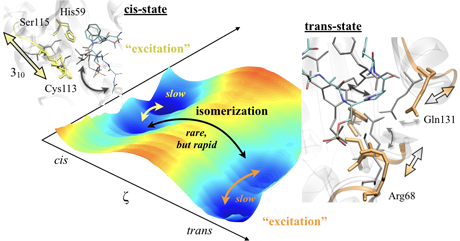
Schematic of enzymatic reaction. Reactions rapidly take place through 'conformational excited states' on a two-dimensional surface expressed by fast and slow variables. (Mori and Sato, J. Pjys. Chem. Lett. 10, 474-480 (2019).)
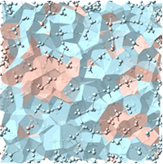

Schematic figure of low- (blue) and high- (red) density local structures in supercooled water. Local density fluctuations generate thermodynamic and dynamic anomalies of water. (Saito et al., J. Chem. Phys. 149, 124504 (8 pages) (2018).)
Selected Publications
- T. Yagasaki and S. Saito, "Fluctuations and Relaxation Dynamics of Liquid Water Revealed by Linear and Nonlinear Spectroscopy," Annu. Rev. Phys. Chem. 64, 55-75 (2013) (invited).
- K. Kim and S. Saito, “Multiple Length and Time Scales of Dynamic Heterogeneities in Model Glass-Forming Liquids: A Systematic Analysis of Multi-Point and Multi-Time Correlations,” J. Chem. Phys. (Special Topic: Glass Transition) 138, 12A506 (13 pages) (2013) (Invited).
- T. Mori and S. Saito, "Conformational excitation and non-equilibrium transition facilitate enzymatic reactions:Application to Pin1 peptidyl-prolyl isomerase," J. Phys. Chem. Lett. 10, 474-480 (2019).
- S. Saito, M. Higashi, and G. R. Fleming, "Site-dependent fluctuations optimize electronic energy transfer in the Fenna-Matthews-Olson protein," J. Phys. Chem. B 123, 9762-9772 (2019).
- J. Ono, Y. Matsumura, T. Mori, and S. Saito, "Conformational dynamics in proteins: Entangled slow fluctuations and nonequilibrium reaction events (Perspective)", J. Phys. Chem. B 128, 20-32 (2024).
- Shinji Saito, "Unraveling the dynamic slowdown in supercooled water: The role ofdynamic disorder in jump motions," J. Chem. Phys. 160, 194506 (13pages) (2024).